 are among the most common causes of systemic mycoses, and because getting a diagnostic assay all the way through FDA clearance is so challenging, it is difficult to incentivize the development of assays that target organisms that may be only rarely, if ever, encountered. 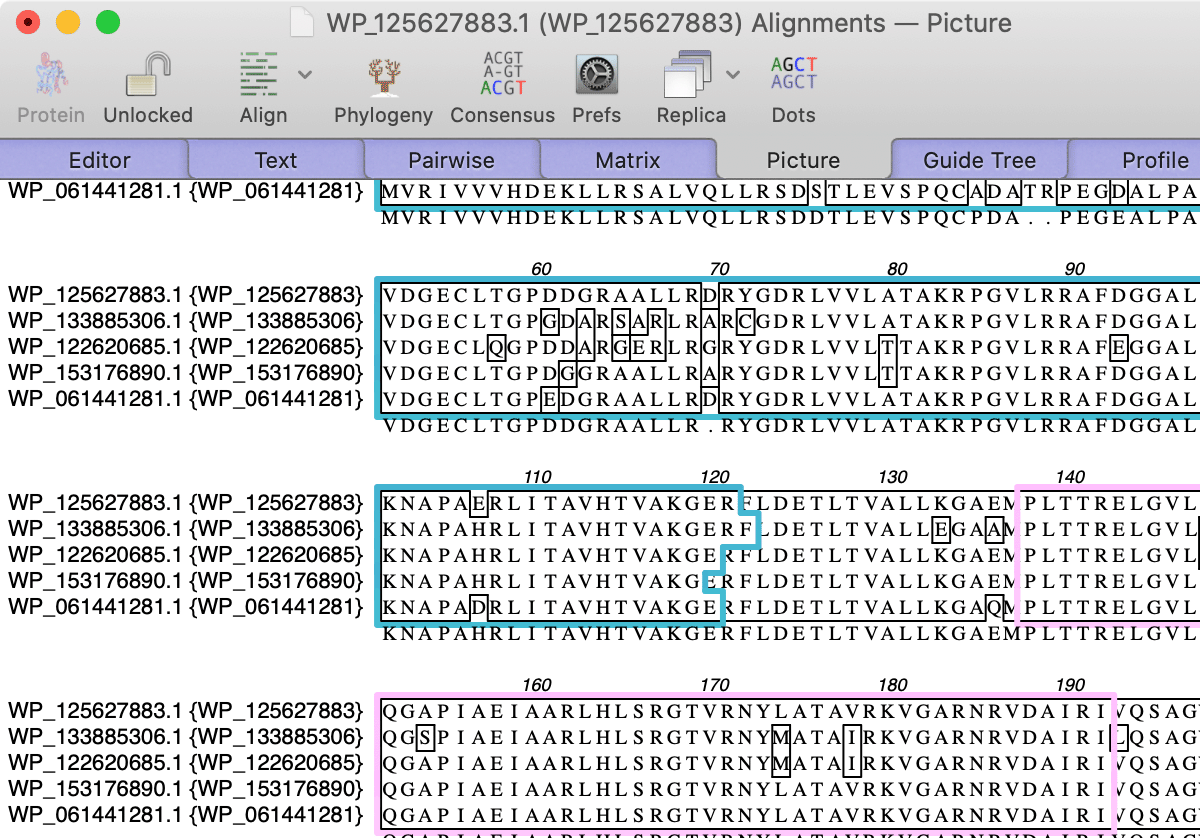
The reasons are both epidemiological and economic. Consequently, virtually all FDA-cleared fungal diagnostic assays focus on a single fungal pathogen or a very limited scope of the most common fungal pathogens ( 12), with most assays directed toward a few Candida species. However, a confounding problem with this need is that because so many human fungal pathogens are so rarely encountered in clinical specimens, it is not possible or financially worthwhile to include them in assays requiring specific analytes to be targeted with specific probes, antibodies, etc., for each potential species. This expansion will necessitate new fungal diagnostic strategies to identify these organisms. Because of the ability of modern medicine to sustain increasingly sicker patients, the continued expansion of the number of fungal species capable of causing infection is to be expected. 
Most of these species are opportunistic pathogens that take advantage of immunosuppressed hosts, with most of them rarely, if ever, seen by the majority of clinical microbiology laboratories. Our own diagnostic program has found more than 1,500 unique species capable of infecting humans ( 12). Since 1995, the number of new fungal pathogens of animals, plants, and humans has increased almost 10-fold ( 11). These results suggest that sequencing the IGS region using nanopore sequencing could be a potential new molecular diagnostic strategy.Īlthough the literature typically cites 300 to 500 species as pathogenic for humans ( 7 – 10), this number is an underestimation, based on case reports alone. Sequencing using the nanopore platform could be completed in less than an hour, and samples could be multiplexed in groups as large as 24 sequences in a single run. neoformans species complexes resulted in only a 74 to 77% identity between the two complexes. Comparison of average percent identities between the C. Phylogenetic analysis of the nanopore- and Sanger-derived sequences resulted in indistinguishable trees. 
Nanopore sequencing errors were predominantly in regions of homopolymers, with G homopolymers displaying the largest number of errors and C homopolymers displaying the least. When the newer R10.3 flow cell was used, accuracy increased to 99.83% identity compared to the same Sanger sequences. Using the R9.4.1 flow cell, IGS sequence identities averaged 99.57% compared to Sanger sequences of the same region. There is enough variation within the two complexes to argue for further resolution into separate species, which we wanted to see if nanopore sequencing could detect. neoformans, which are the main pathogenic members of this genus, using the Oxford Nanopore Technologies MinION device and Sanger sequencing. We sequenced isolates from two Cryptococcus species complexes, C. We investigated the fungal intergenic spacer (IGS) sequence in combination with nanopore sequencing for fungal identification. Fungal infections are being caused by a broadening spectrum of fungi, yet in many cases, identification to the species level is required for proper antifungal selection. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
